Formation of repressive complex in the mammalian circadian clock is demoted by the secondary pocket of CRY1
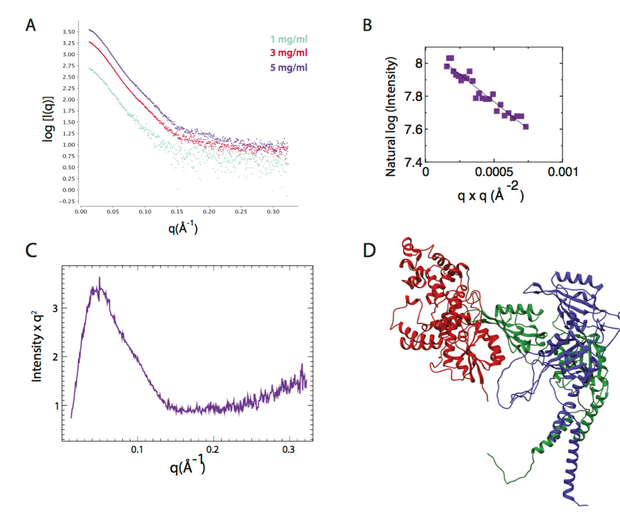
In this paper, the authors show that CRY1, a protein coding gene that activates circadian gene expression and metabolic states and circadian oscillators, binds directly to the PAS domain core of CLOCK:BMAL1. Precise control of CLOCK:BMAL1 activity by coactivators and repressors establishes the ~24 hr periodicity of gene expression. Integrative modeling and solution X-ray scattering…
Read more