Two recent papers from Doudna and Marletta groups combine SAXS with Cryo-EM
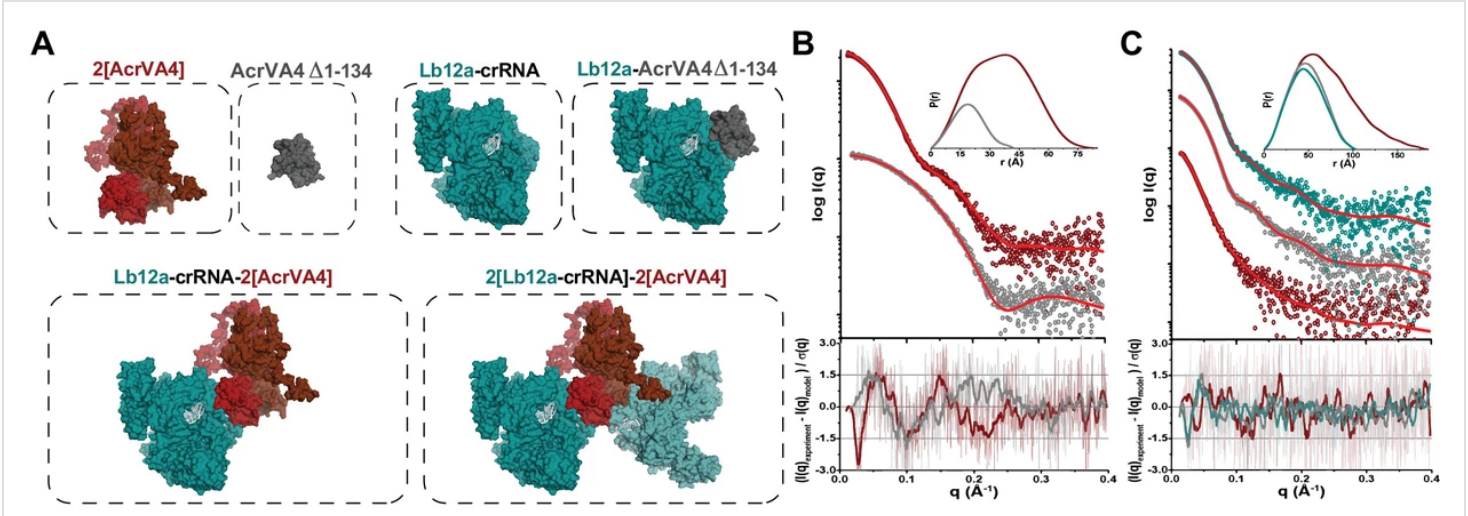
The Doudna group, explained in their recent paper, “Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a” combine SAXS with Cryo-Em to show the C-terminal binding domain is sufficient for Cas12a inhibition. CRISPR-Cas systems provide bacteria and archaea with programmable immunity against mobile genetic elements. Evolutionary pressure by CRISPR-Cas has driven bacteriophage to evolve small protein inhibitors, anti-CRISPRs (Acrs), that block Cas enzyme function by wide-ranging mechanisms. The inhibitor AcrVA4 uses a previously undescribed strategy to recognize the L. bacterium Cas12a (LbCas12a) pre-crRNA processing nuclease, forming a Cas12a dimer, and allosterically inhibiting DNA binding. Their findings explain a new mode of CRISPR-Cas inhibition and illustrate how structural variability in Cas effectors can drive opportunistic co-evolution of inhibitors by bacteriophage.


In the Marletta group’s recent paper, “Allosteric activation of the nitric oxide receptor soluble guanylate cyclase mapped by cryo-electron microscopy” they corroborate the structural elongation of the particle observed in cryo-EM using small angle X-ray scattering (SAXS). These structures delineate the endpoints of the allosteric transition responsible for the major cyclic GMP-dependent physiological effects of NO. Soluble guanylate cyclase (sGC) is the primary receptor for nitric oxide (NO) in mammalian nitric oxide signaling. They determined structures of full-length Manduca sexta sGC in both inactive and active states using cryo-electron microscopy. NO and the sGC-specific stimulator YC-1 induce a 71° rotation of the heme-binding β H-NOX and PAS domains. Repositioning of the β H-NOX domain leads to a straightening of the coiled-coil domains, which, in turn, use the motion to move the catalytic domains into an active conformation. YC-1 binds directly between the β H-NOX domain and the two CC domains.